DynamiR
Welcome to DynamiR
🧬 What This Tool Does
DynamiR is a desktop application designed for researchers and biologists to identify and analyze potential binding sites where microRNAs can attach to messenger RNA (mRNA) sequences.
Key Capabilities:
- Direct Analysis (Matching Mode): Analyze a specific microRNA against a target mRNA to find exact binding positions
- Database Scanning (Dynamite Mode): Scan all ~48,860 known microRNAs against your target mRNA in one analysis
- Multiple Input Methods: Load sequences from files (FASTA, GenBank) or directly from online databases (miRBase, NCBI)
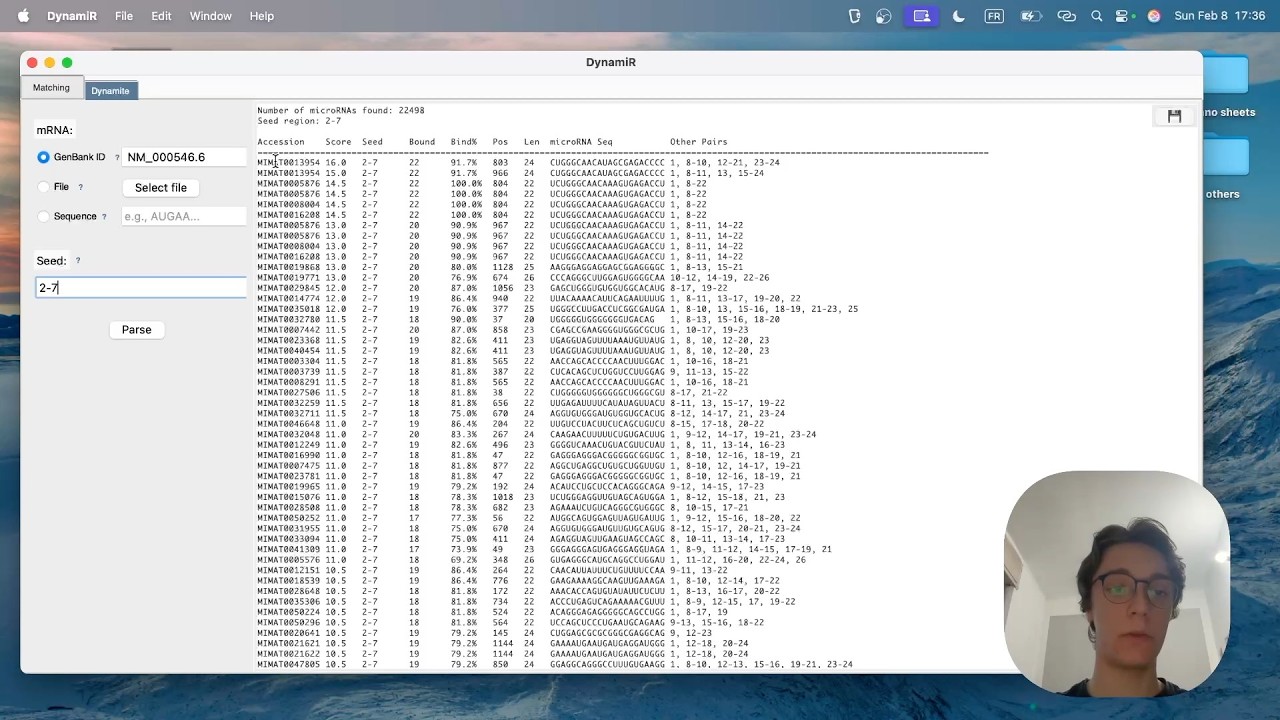
- Results: Get detailed binding site information including positions, pairing statistics, and alignment visualizations
- Export Functionality: Save your analysis results for further processing or sharing
⚡ Quick Start
Download & Installation
- 📥 Download for macOS - Version 1.0 - Available
First Steps
- Install the application for your operating system
- Launch the application
- Choose a Mode - This step can be done in two ways:
- Use Matching Mode to test a specific microRNA
- Use Dynamite Mode to discover all possible microRNA binders
- Enter Your Data: Provide sequences via files or database IDs
- Run Analysis and view results
- Export your findings
� Video Demonstration
Watch this video to see DynamiR in action:
Full Demo on YouTube - See how to use all analysis modes and interpret results.
�🎯 Use Cases
Research & Discovery
- Identify potential microRNA regulators for your target genes
- Validate predicted binding sites computationally
- Discover off-target microRNA interactions
Validation
- Confirm microRNA-mRNA interactions predicted by other tools
- Test multiple seed regions to understand binding specificity
- Compare results across different analysis parameters
Educational
- Learn how microRNA binding prediction works
- Understand seed-region importance in sequence matching
- Visualize RNA duplex alignments
📚 Documentation
- Installation Guide - Setup instructions for all platforms
- Getting Started - First time users guide
- Matching Mode - Direct microRNA-mRNA analysis
- Dynamite Mode - Database-wide scanning
- Biological Concepts - Understanding the biology
- How It Works - System architecture and workflows
💡 Key Features
✅ Dual Analysis Modes - Choose between focused analysis or broad screening
✅ Multiple Input Options - Files, database IDs, or direct sequence entry
✅ Visual Alignments - See base pairing details at each binding site
✅ Quick Export - Save results in text format
✅ Responsive Interface - User-friendly experience
✅ Quality - Tool with error handling
🔬 What Makes This Different
This tool implements seed-based matching with Watson-Crick pairing detection and wobble pair recognition. It uses proper bioinformatic algorithms to identify realistic binding scenarios.
Seed Region Matching: The default seed region is positions 2-7 of the microRNA - this is the critical region for binding stability. You can customize the seed region to match your research needs.
Extended Analysis: Full duplex alignment scoring shows additional pairing strength beyond the seed region.
📖 Learn More
- New to microRNA biology? Start with Biological Concepts
- Want to understand the matching algorithm? Read How It Works
- Ready to analyze? Follow Getting Started
🤝 Support
For questions, issues, or suggestions: - 📧 Email: remi.hocquetmartin@icloud.com - 🐛 Report bugs: GitHub Issues
Version: 1.0
Last Updated: November 2025
License: © 2025 HOCQUET MARTIN Rémi